Biography
Interests
Fardous El-Senduny, F.1,3, Farid Badria, A.2, Ahmed EL-Waseef, M.3, Eduardo Callegari, A.4 & Fathi Halaweish1*
1Chemistry and Biochemistry Department, South Dakota State University, Brookings, USA
2Pharmacognosy Department, Faculty of Pharmacy, Mansoura University, Egypt
3Chemistry Department, Faculty of Science, Mansoura University, Egypt
4Division of Basic Biomedical Sciences, Sanford School of Medicine, The University of South Dakota, USA
*Correspondence to: Dr. Fathi Halaweish, Chemistry and Biochemistry Department, South Dakota State University, Brookings, USA.
Copyright © 2018 Fathi Halaweish, et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Abstract
The first line in chemotherapy for ovarian cancer is platinum-based drugs. Despite combination treatment regimes, the 5-year survival rate still in decrease due to the resistance of the recurrent disease to the treatment. Finding a natural product that can restore cell’ sensitivity to the common therapeutic drugs is needed. Recent finding in our research showed that cucurbitacin B sensitizes the cisplatin-resistant ovarian cancer cell lines to the cytotoxicity of cisplatin.
The aim of this work is to investigate the mechanism behind the sensitization by cucurbitacin B and analyzing change in protein expression after treatment with either cucurbitacin B, cisplatin or combination of both compounds. The protein lysate was subjected to 2D-nano-UPLC-MS/MS analysis. Data analysis showed that cucurbitacin B as a single compound or in combination with cisplatin treatment mainly altered proteins involved in fatty acid synthesis, antioxidant system, protein folding, cytoskeleton, DNA replication, translation and chromatin assembly. Western blot validation for FASN, LPPRC and H2A.x was in agreement with the proteomic data confirming the possible mechanism of sensitization effect of cucurbitacin B via the change in proteins involved in the induction of apoptosis and inhibition of survival pathways.
Introduction
Ovarian cancer is the fifth leading cause to the cancer-related mortality in women. In 2016, there will be
22,280 new cases of ovarian cancer diagnosed and 14,240 women will die of ovarian cancer in the United
States. The five-year survival rate is 46.2% and the mortality rates for ovarian cancer have declined only
slightly in the forty years [1,2]. This short-term survival is due to the lack of the early detection methods.
The first line of chemotherapy treatment is platinum-based drugs such as cisplatin and carboplatin. Most
patients suffer from cancer recurrence and became resistant to the chemotherapy regimen. The resistance is
due to the increase in drug efflux, detoxification of the drug, high tolerance to sub lethal DNA damage or
the capability to repair DNA damage [3]. Till now, research still search for new key in resistance of ovarian
cancer to chemotherapy, which will guide the selection of specific inhibitor or combination therapy.
Cucurbitacins are highly oxidized tetracyclic triterpenoid. Cucurbitacin B and D are most dominant types in the plant followed by cucurbitacin E, G, H and I. They are used in Chinese remedies for hepatitis, inflammation and cancer treatment. Cucurbitacin B is the most studied one followed by cucurbitacin E [4-6]. Cucurbitacin B shows anti-proliferative activity against wide range of cancer cell lines such as breast, lung, glioblastoma, hepatic, pancreatic, colon, central nervous system, oral, ovarian and leukemia [7-17]. Moreover, cucurbitacin B increased the cytotoxicity of different chemotherapeutic agents including methotrexate toward osteosarcoma [18], cisplatin against cutaneous squamous cell carcinoma cell lines and laryngeal squamous cell carcinoma cells [19,20], gemcitabine against pancreatic cancer [21,22] and docetaxel in laryngeal cancer [23]. Also cucurbitacin B sensitized the breast cancer cells to the radiotherapy [24]. This sensitization effect of cucurbitacin B attributed to the cell cycle arrest, cytoskeleton alteration [16,25,26], inhibition of signal transducer and activator transcription 3 (STAT-3) [17,18,20,22,27], inactivation of mitogen activated protein kinase pathway [7]and reducing the glutathione content [28]. Recently, we have showed that cucurbitacin B is able to restore the sensitivity of cisplatin-resistant ovarian cancer cells (A2780CP) to cisplatin cytotoxicity [9]. This sensitization effect of cucurbitacin B was found to be due to the inhibition of ERK and STAT-3 phosphorylation, a decrease in total glutathione, an increase in reactive oxygen species and via a decrease in DYRK1B (dual-specificity tyrosine-regulated kinase). Nowadays, a single drug treatment of ovarian cancer is less favorable because of the increase in resistance against its cytotoxicity and using cisplatin combined with other chemotherapeutic drugs is more effective than single drug therapy to destroy the tumor by targeting different pathways such as cisplatin combined with paclitaxel in the treatment of ovarian cancer.
It has been proven that proteomic analysis can be used for drug discovery and identification of targets affected by the treatment and may open doors for a novel proteins that may play a role in the cancer metastasis or treatment [29]. To our knowledge, there is only one report that showed the proteins expressed upon cucurbitacin B treatment of breast cancer cell line (MDA-MB-231 and MCF-7) [25]. Duangmano et al. [25] studied the proteomic change by two- dimensional gel electrophoresis and were able to detect only four differentially expressed proteins. Cucurbitacin B down regulated nucleophosmin/B32 and up regulated three heat shock-related proteins (HSP70, prolyl 4-hydroxylase and chaperonin). In this article, we investigated the change in protein expression after cucurbitacin B alone or combined with cisplatin treatment for cisplatin-resistant ovarian cancer cell line (A2780CP) by using 2D-nano-UPLC-MS/MS.
Results and Discussion
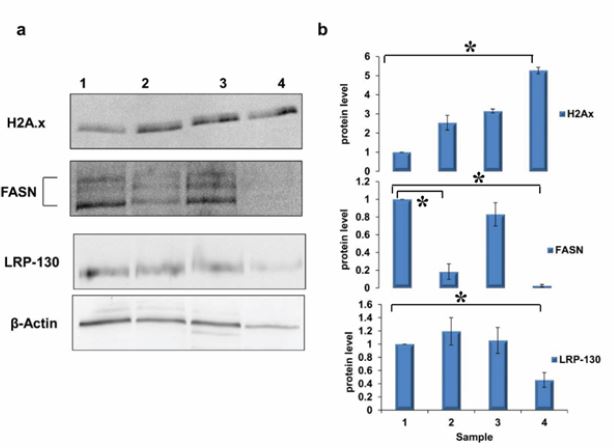
Recent study in our group documented [30] the ability of cucurbitacin B to sensitize the cisplatin-resistant
ovarian cancer cells (A2780CP) in both 2D and 3D culture to the cytotoxicity of cisplatin. The proteomics
analysis showed that the combination treatment down regulated the protein level of Leucine-rich PPR motifcontaining
protein (LRP130) (Figure 1, Supplementary material table S1). LRP130 contains 11 pentatrico
peptide repeats which binds to the hydrophilic nucleotides [31]. It plays a role in the cellular homeostasis
and cytoskeleton organization [23,33] through binding to mitochondrial and nuclear RNA controlling
the RNA transport machinery [34]. Moreover, it has been reported that its transactivation activity in the
multidrug resistance genes such as p-glycoprotein MDR1 [35] confer the antiapoptotic properties to the
cancer and decrease the translocation of the drug into the cancer cells. This is the first report to demonstrate
the inhibitory effect of either cucurbitacin B or cisplatin on the expression level of LRP130. This effect may
lead to an increase in the amount of cisplatin in the cells and increased response of the resistant ovarian
cancer cells to the cytotoxicity of cisplatin.
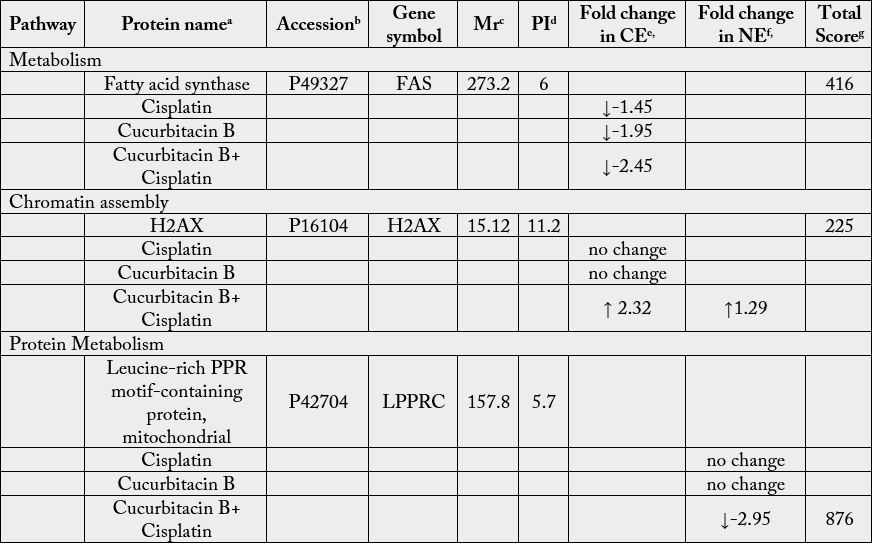
aThe protein name corresponding to the entry at the Swiss Protein database.
bFunction annotations were retrieved from Swiss Protein (http://www.uniprot.org/) and NCBInr (http://www.ncbi.
nlm.nih.gov/) for human (Homo sapiens)
cThe theoretical molecular mass or molecular weight (MW) coming from database of the protein identified.
dThe theoretical isoelectric point (pI) coming from database of the protein identified.
e and fThe ratio shows the expression or relative abundance of the protein through the heat map analysis by comparing
spectral counts, peptides and percentage of coverage followed by NSAF (Normalized Spectral Abundance Factor),
using a “label free” approach. The values are represented by the log2 relative expression between the replicates of the
biological samples of each group. Ratios e and fcorrespond to treated cells/control in cytoplasmic extract or nuclear extract
(other cellular components except cytoplasm), respectively.
gScores greater than 40 were taken as significant match
based on MOWSE score (http://www.matrixscience.com), the threshold was set up by the server at the significance
level P≥0.05 for random hit.
hthe molecular function of each protein was retrieved from BANTHER website (http://www.pantherdb.org/).
Here in this study, the treatment of A2780CP cells with cucurbitacin B and cisplatin together significantly down regulated the level of FASN (Figure 1, Supplementary material table S1). Cucurbitacin B alone decreased the level of FASN which is a further indication of cucurbitacin B as a potential chemosensitizer for cisplatin chemotherapy (Figure 1, Supplementary material table S1). FASN plays a major role in the lipogenesis. It is considered as an oncogene in breast and prostate cancer [36,37] by stabilizing β-catenin in the cytoplasm and activation of Wnt-1/β-catenin pathway [38]. Moreover, the inhibition of fatty acid synthase activity by RNAi in osteosarcoma inhibited the cell invasion and metastasis via the inhibition of HER2/PI3K/AKT pathway [39,40]. Furthermore, its down regulation by C-75, a selective FASN inhibitor, potentiated the cytotoxicity of cisplatin in ovarian cancer via induction of apoptosis [41]. Treating the cells with cucurbitacin B and cisplatin together led to an increase in the free H2A variants in the cytoplasm (Figure 1, Supplementary material table S1) leading to genome instability and increasing the sensitivity of the cells to DNA damaging by cisplatin. The free H2Ax in the cytoplasm was found to be increased in Imatinib-treated gastrointestinal stromal tumors (GIST) a hallmark for the apoptosis induction after Erlotinib treatment [42-44].
Experimental
The cisplatin-sensitive (A2780) and cisplatin-resistant (A2780CP) ovarian cancer cell lines [31] were
obtained from Dr. Stephen Howell, University of California, San Diego, USA. The cells were maintained
in RPMI-1640 (Thermoscientific™ Hyclone™) supplemented with 10% fetal bovine serum (FBS)
(Thermoscientific™ Hyclone™) and 1% penicillin (100IU/mL)/streptomycin (100μg/mL) (Corning™
Cellgro™). In order to maintain the resistance, 1μM cisplatin was added to the medium every 2-3 passages.
Cisplatin (cis-diamineplatinum (II) dichloride) was obtained from Sigma and dissolved in 0.9% saline then
stored as 8mM stock solution at -20°C. Cucurbitacin B was isolated from Cucurbita texana and characterized
by spectroscopic techniques in our lab [46].
A2780CP cells were seeded in 25cm2 culture flask and incubated overnight. The cells were incubated either
with 2μM cucurbitacin B, 40μM cisplatin for 48hrs or pre-incubated with 2μM cucurbitacin B for 24hrs then incubated for another 48hrs with 40μM cisplatin. Cell fractionation was done to increase the number
of identified proteins in each sample. After cisplatin-resistant ovarian cancer cells were treated, the cells were
washed with ice-cold 1x PBS and collected after trypsinzation. The cell pellet was washed with 1x PBS and
re-suspended in low salt buffer (10mM HEPES pH: 7.9, 10mM KCl, 1.5mM MgCl2.6 H2O) and incubated
20min on ice. The cells were homogenized using 20 strokes by dounce homogenizer. The cell lysate was
centrifuged at 800 × g at 4°C. The supernatant constituted a fraction enriched in cytosol components was
transferred to a pre-chilled tube and lysed by 1x lysis buffer (50mM Tris-HCl pH: 7.5, 150mM NaCl, 1%
Triton X-100, 50mM NaF, 1mM DTT, without protease inhibitor cocktail). The pellet enriched in the rest
of the components of the cells, mainly membranes and organelles, were lysate using lysis buffer. The lysate
was centrifuged at 13,000rpm for 30min, and the supernatant transferred to a new pre-chilled tube. Protein
concentration was estimated by BCA protein assay kit, Pierce™.
The proteins obtained from cell lysate (individual cytoplasmic and nuclear fractions) were in-solution
reduced with DTT (Sigma Aldrich, St. Louis, MI) at 65°C for 5 minutes, followed by alkylation with
Iodoacetamide (Sigma Aldrich, St. Louis, MI), and digested with sequencing grade trypsin (Promega,
Madison WI) overnight at 37°C. The digested peptides were concentrated by vacuum centrifuge (Speed
Vac, ThermoScientific). The trypsin digested peptides were dissolved in 100mM Ammonium Formate, pH
10, and separated through 2D-nanoLC with dilution using a 2D-nanoAcquity UPLC (Waters, Milford,
MA) according to Dong G et al. [47,48].
The first dimension was performed in XBridge BEH130 C18, 5μm, 300μm×50mm NanoEase Column (Waters, Milford, MA) using solvent A1: 20mM Ammonium Formate, pH=10 and B1: 100% Acetonitrile (Fisher Optima, LC-MS grade). The flow at 1st dimension was 2μL/min and 10 different step gradients (dilution method) were performed for 20 minute separately. The second dimension included trapping and desalting online through 180μm×20mm, 5μm Symmetry C18 nanoAcquity UPLC trap column (Waters, Milford, MA) at flow = 20μL/min, 99% A2 (H2O, 0.1% Formic Acid) and 1% B2 (100% Acetonitrile, 0.1% Formic Acid) for 20 minutes. After the peptides were desalted and concentrated, they were separated online in the second dimension through BEH130 C18 1.7um, 100 μm×100 mm nanoAcquity UPLC column (Waters, Milford, MA). The standard solvent gradient used was: 0-2min, 3% B2 isocratic; 2-40min, 3-85% B2 linear, at a flow rate of 400nL/min for 60 minutes. The eluted ions were analyzed by one full precursor MS scan (400-150m/z) followed by four MS/MS scans of the most abundant ions detected in the precursor MS scan while operating under dynamic exclusion or direct data acquisition system (Table 1). Spectra obtained in the positive ion mode with nano ESI-Q-Tof Synapt G1 mass spectrometer (Waters, Milford, MA) were deconvoluted, and analyzed using the MassLynx software 4.1 (Micromass, UK), and exported to ProteinLynx Global Server v3.0 (PLGS v3.0) (Waters, Milford, MA) to creates the list of masses. A peak list (PKL format) was generated to identify +1 or multiple charged precursor ions from the mass spectrometry data file (Table S2). The instrument was calibrated in MS/MS mode using 100 fmole of (Glu1) - Fibrinopeptide B human (Sigma, Saint Louis, MO) with a RMS residual of 3.857 e-4 amu or 6.9413 e-1 ppm. Parent mass (MS) and fragment mass (MS/MS) peak ranges were 400-2000 Da and 65- 2000 Da, respectively.
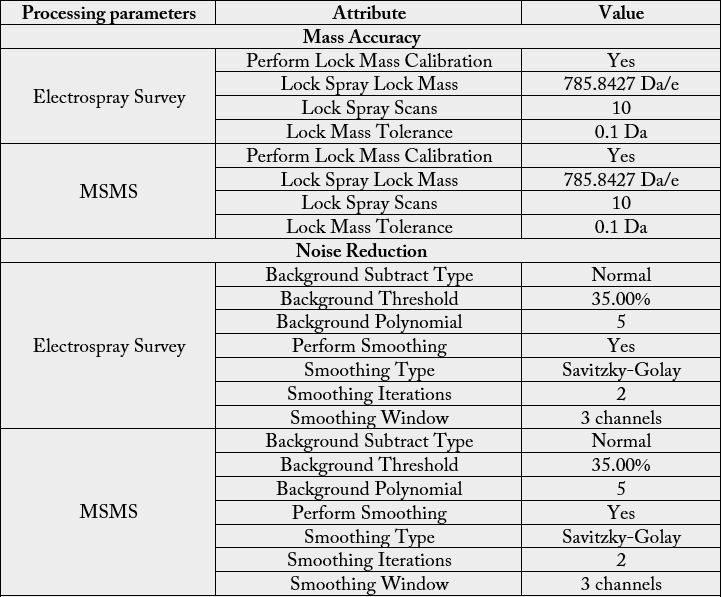
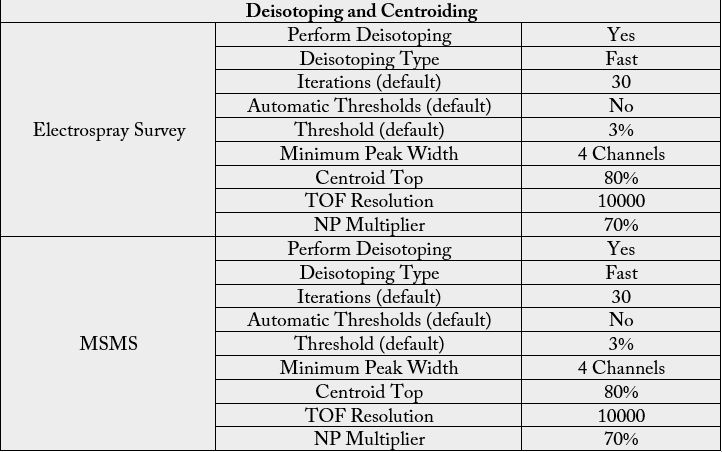
Mascot server v2.5.0 and Mascot Daemon Toolbox v2.5.0 (www.matrix-science.com, UK) in MS/MS
ion search mode (local licenses) were applied to conduct peptide matches (peptide masses and sequence
tags) and protein searches against Swiss Protein 2015_03 (547964 sequences; 195174196 residues) using
taxonomy filter Homo sapiens (humans) (20203 sequences). The following parameters were set for the
search: carbamidomethyl (C) on cysteine was set as fixed; variable modifications included asparagine and
glutamine deamination and methionine oxidation, as well as Error Tolerance mode. One missed cleavage
was allowed for regular searching; monoisotopic masses were counted; the precursor peptide mass tolerance
was set at 50ppm; fragment mass tolerance was 0.3 Da and the ion score or expected cut-off was set at 5.
The MS/MS spectra were searched with MASCOT using a 95% confidence interval (C.I. %) threshold (p
< 0.05), with which minimum score of 40 was used for peptide identification. All proteins identified were
found these domains. Also, they were corroborated with X! Tandem (GPM) (www.thegpm.org), an open
source setting using similar setup like Mascot (enzyme, cleavages, and modifications) and ENSEMBL
Homo sapiens database (http://ftp.ensembl.org/pub/release-79/embl/homo_sapiens/).
The protein relative abundance analysis was performed using a “label free” approach through the measure of the spectral counts and the ratio reported as the log2 relative expression. ProteoIQ software v2.7 (www. premierbiosoft.com) was used for the post-mass spectrometry bioinformatics analysis (ratio calculation and normalization by NSAF=Normalized Spectral Abundance Factor, as well as the log2 relative expression) mentioned before. The proteins accepted from the identification list met the following criteria: >99.0% probability to achieve a false discovery rate (FDR) of < 1% [49,50] (the analysis was performed using SP with taxonomy filter for human target and decoy databases at ProteoIQ v2.7). Moreover, the proteins identified were corroborate by the observation of the False Positive Rate (FPR) coming from the analysis by GPM as second approach (the FPR average was 0.62%) [51,52].
The differentially expressed proteins were submitted to PANTHER database to identify the molecular
function, biological process and pathways of the proteins (http://www.pantherdb.org/). Proteins that are
involved in the survival of cancer patients were identified by using Bioprofiling website (http://www.
bioprofiling.de/GEO/GENESURV/genesurv.html). GPM server was used to identify the enrichment and
depleted proteins in the protein list.
The cisplatin-resistant ovarian cancer cells were seeded in 60 mm culture dish (5*104cells/mL, 3mL/dish)
and allowed to adhere overnight at 37°C and 5% CO2. The cells were incubated either with 2μM cucurbitacin
B, 40μM cisplatin for 48hrs or pre-incubated with 2μM cucurbitacin B for 24hrs then further incubated for
another 48hrs with 40μM cisplatin. The cells were washed 2 times with 1x PBS and lysed by 1x RIPA buffer
containing 1x protease and phosphatase inhibitor cocktail. The protein content was quantified by BCA
protein assay kit. 20 or 40μg/well was separated on SDS-PAGE.
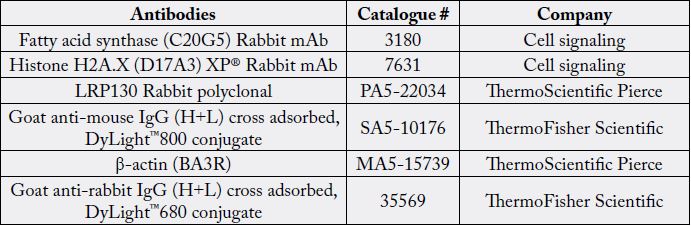
The protein was transferred to a nitrocellulose membrane 0.2 or 0.45μm for 45 or 90minutes at 100 or 90 V, respectively. The membrane was reacted with the primary antibodies (Table S3). β-actin was used as a loading control. The secondary antibody (Table S3) was applied to the membrane and was detected either with the GE Healthcare Amersham ECL Plus Western Blotting Detection Reagents (Piscataway, NJ) and the signal was recorded by a UVP gel documentation system (UVP, Upland, CA) or by Odyssey® Imager. The level of protein expression was normalized to the level of the loading control β-actin then the fold of change was determined by comparing the band intensity of the control cells to the untreated cells. Statistical analysis was done using the software SPSS 19 for Microsoft Windows (SPSS Inc, Chicago, Illinois). The fold of change in Fatty acid synthase, LRP-130 and Histone H2A.X level was tested using one-way analysis of variance (ANOVA). The fold of change was considered significant when (P < 0.05).
Conclusion
Overcoming the ovarian cancer resistance will led to a better survival rate of the patients. Using natural products such as cucurbitacin B at low concentration with the common chemotherapeutic drug, will improve the cytotoxicity of the drug with minimal side effects. In our report, cucurbitacin B alone was able to alter the expression of vital proteins required to regulate different pathways in the tumor cells. When cucurbitacin B combined with cisplatin, they efficiently kill the tumor cells by changing the expression of wide range of proteins that control survival signaling pathways.
Here we show that this combination worth further investigation on the level of in vivo studies followed by a clinical trial.
Supplementary Files
The procedures for all assays including cell treatment and fractionation for 2D-nanoLC, protein identification
by 2 dimensional nano-liquid chromatography tandem mass spectrometry (2D-nano-UPLC-MS/MS)
analysis, database searching, gene ontology annotation and western blot confirmation for proteomic results
are reported in the supplementary file.
Acknowledgements
This work was supported by the Egyptian government via the Egyptian Ministry of Higher Education
and Scientific Research and was supported by the National Institute of General Medical Sciences of the
National Institutes of Health under grant number P20GM103443 and P20RR016479.
Bibliography

Hi!
We're here to answer your questions!
Send us a message via Whatsapp, and we'll reply the moment we're available!