Biography
Interests
Amr Saeb, T. M.1, Shatha Makki, M.2 & Hamsa Tayeb, T.3,4*
1Strategic Center for Diabetes Research, College of Medicine, King Saud University, Saudi Arabia
2King Faisal Specialist Hospital International Holding Co, Saudi Arabia
3Department of Genetics, King Faisal Specialist Hospital and Research Center, Riyadh, Saudi Arabia
4Saudi Human Genome Project, King Abdulaziz City for Science and Technology (KACST), Riyadh, Saudi Arabia
*Correspondence to: Dr. Hamsa Tayeb, T., Department of Genetics, King Faisal Specialist Hospital and Research Center & Saudi Human Genome Project, King Abdulaziz City for Science and Technology (KACST) Riyadh Saudi Arabia.
Copyright © 2020 Dr. Hamsa Tayeb, T., et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Introduction
The current novel coronavirus (COVID-19) outbreak is regarded as a crucial public health emergency of
international concern (PHEIC). It is involving a novel type of coronavirus initially identified in Wuhan,
China. At the time of writing, there have been 167511 confirmed cases around the world with 6606 deaths.
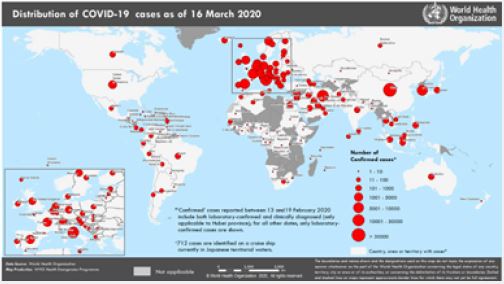
In Saudi Arabia, there are 103 confirmed cases classified as Local transmissions [1] Figure (1). The global
risk assessment was set as very high for China, regional and global levels.
Based on a recent meta-analysis of symptoms in 50,466 COVID-19 patients, the most common symptoms of severe COVID-19 infections are fever and cough, 89.1% and 72.2%, respectively [2]. Additionally, myalgia or fatigue (11-52%), and shortness of breath (3-31%) occurs in patients at illness onset. The fatality rate was estimated to be 4.3% and the majority of fatalities occurred amongst adults (median age 59 years) [3]. Older patients and those with chronic medical conditions might be at higher risk of the disease, however, the US Centers for Disease Control and Prevention (CDC) interim clinical guidance for risk factors of the severe illness is not clear [4]. The causal agent of Covid-19 is severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) that belongs to family Coronaviridae. The family Coronaviridae encompasses A group of large, single, plus-stranded RNA viruses isolated from several animal species and common cold and diarrheal illnesses in humans are widely recognized [5,6]. In China, and in 2003, the latest strain of coronavirus (severe acute respiratory disease) syndrome The SARS outbreak was associated with the [SARS-CoV] coronavirus [5,6]. In Saudi Arabia, MERS-coronavirus (MERS-CoV) was published in 2012. It is the causal agent of the Middle East respiratory syndrome (MERS) that is also known as camel flu. This virus causes respiratory infection with mild to severe symptoms that typically include fever, cough, diarrhea, and breathing difficulties. Elder patients and those with compromised immunity usually exhibit more severe symptoms [7,8]. The new SARS-CoV-2 belongs to the Orthocoronavirinae subfamily, is distinct from MERS‐CoV and SARSCoV [9]. The zoonotic transmission was suggested as the source of the epidemic since numerous patients confirmed the visitation of a local fish and wild animal market in Wuhan during November. Besides, human to human transmission was also reported [10,11]. No doubt gaining more information about molecular epidemiology, evolutionary models, and phylogenetic analysis can aid in estimating genetic variability, the evolutionary rates, genetic mutations and divergence which in turn have important consequences for disease progression, virus evolution, and drug and vaccine development.
Literature Review
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2); formerly known as 2019-nCoV was
reported in Wuhan (COVID-19), a city in the Hubei Province of China, as the causal agent of a cluster of
pneumonia cases. n November 2019 [1-6].
Since December 2019, different cases of pneumonia of unknown origin associated with perpetuity at the
Wuhan market in China have been reported. To date, more than 81077 cases were reported in China (Hubei
Province). Collaborative investigations should that the epidemic peaked in late January to the beginning
of February 2020 in China [12]. Though the majority of reported cases are from Hubei and surrounding
provinces, many other cases have been reported in other cities and provinces throughout mainland China
[13,14]. Travelers from China then individuals who interacted with them caused the increase of cases
in other countries across all continents, with Antarctica as an exception, and with a proportional rate of
transmission [1,15-17]. However, local transmission has led to lesser outbreaks in other countries. In Italy,
local transmission resulted in 24747 confirmed cases of infection accompanied by 1809 total death cases.
Besides, in Iran (the Islamic Republic of) local transmission resulted in 14991 confirmed cases of infection
accompanied by 853 total death cases. Moreover, in the Republic of Korea local transmission resulted in
8236 confirmed cases of infection accompanied by 75 total death cases [1]. Furthermore, local transmission
was the main cause of the infection in the Gulf states resulting in 401 cases in Qatar, 221 in Bahrain, 112
in Kuwait, 98 in the United Arab Emirates and 22 infections in Oman. Upon writing this report and
based on the latest WHO situation report released 10 AM CET 17 March 2020, Saudi Arabia reported
133 confirmed infection cases and 30 total new cases all by the method of local transmission [18]. Though
we lack a complete understanding of the transmission risk, epidemiological investigations in Wuhan city
showed that the outbreak started in a seafood market that also traded live wild animals, in which the majority
of the patients had worked or visited. However, this marketplace was later on shut down and disinfected
[19]. Late on, person-to-person transmission became the main mode of transmission. The main thought of
a way of SARS-CoV-2 spread from person-to-person is via respiratory droplets. Virus particles are released
with respiratory secretions Person coughs, sneezes, or speeches when contaminated. The virus can establish
infection in the next person when it reaches his/her mucous membrane directly or indirectly (by touching an
infected surface) through eyes, nose or mouth. Thus the COVID-19 virus is transmitted between individuals
through close contact and droplets, not by airborne transmission, based on the available evidence [20].
However, airborne transmission should not be completely disproved in the time being [21,22]. It was
suggested that transmission is more likely soon after the onset of symptoms since the Viral RNA levels
appear to be higher in this stage compared with the final stages of the disease [17], however, more research
is needed to validate this hypothesis. The rates of transmission differ in different reports. Secondary infection
rate was 1-5% in China and 0.45% in USA [23,24]. Several reports showed that asymptomatic individuals
and/or individuals within the incubation period can transmit SARS-CoV-2 [25-29]. Even though the live
viruses were cultivated from stools [30], and fecal-oral transmission did not appear to be an essential factor
in the spread of infection [23].
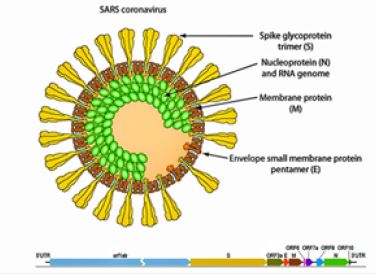
• Taxonomy: Category IV ((+) ssRNA); Coronaviridae; Coronavirinae; Betacoronavirus; Sarbecovirus;
Severe acute respiratory syndrome coronavirus 2
• Virion: enveloped, spherical, with a diameter of 60-140nm with spikes of around 9-12nm
• Genome: ~30 kb positive-sense, RNA single-stranded
• Transcript of RNA: 5 'hat, 3' poly-A tail
• Proteome: 10 proteins
• Transmission: Some early cases had some link to the market for seafood and livestock, indicating
transmission between animals and humans. Later cases suggest continuous transmission from human to
human.
Phylogeny: More closely aligned with bat-SL-CoVZC45 and bat-SL-CoVZXC21 than with SARS-SLCoVZXC21.
• Epidemiology: first published in December 2019 in Wuhan, China. The virus has spread rapidly to other
areas of China and other nations, including
• Clinical: 2019-nCoV cases reported have varied from no symptoms to serious pneumonia and death. There
may be signs such as fever, cough, shortness of breath.
• References: GenBank: MN908947; PMID: 31978945; CDC - 2019-nCoV
• Other Names: 2019-nCoV
Full-length genome sequence of SARS-CoV-2 is almost identical and share 79.6% sequence identity to SARS-CoV. Moreover, SARS-CoV-2
At the whole-genome stage, 96 per cent is the same as a bat coronavirus. Analysis of the pairwise protein sequence of seven domains of non-structural proteins indicates that this virus belongs to the species of SARSr-CoV. In addition, SARS-CoV-2 uses the same cell entry receptor-angiotensin converting enzyme II (ACE2)-as SARS-CoV. Full-genome sequencing and phylogenic analysis indicated that the coronavirus that causes COVID-19 is a betacoronavirus in the same subgenus as the severe acute respiratory syndrome (SARS) virus (as well as several bat coronaviruses), but in a different clade [31]. It was found that MERSCoV is distantly related to the new virus [31,32]. The SARS-CoV-2 genome, it is 75 to 80 percent the same as SARS-CoV and is much more closely linked to other bat coronaviruses.
And it appears likely that bats are the primary source; whether The SARS-CoV-2 is transmitted directly from bats or through some other mechanism is unknown [33]. Population genetic analyses of 103 genomes of SARS-CoV-2 indicated that these viruses formed into two major groups, called L and S, well characterized by two distinct SNPs. Although the L type (about 70 percent) is more prevalent than the S type (about 30 percent), the ancestral variant was found to be the S type. Although the L type was more prevalent in the early stages of the Wuhan outbreak, the L type incidence decreased after the beginning of January 2020. More extreme selective pres may have been imposed by human engagement which could be more violent and quicker to spread. The S type, which is on the other hand, evolutionarily older and less aggressive, due to comparatively weaker selective pressure [34], it may have increased in relative frequency. Coronaviruses are naturally hosted by bats and are formed evolutionarily. Receptor, genetic evolution.
Binding and pathogenesis revealed that through sequential recombination of bat SARSr-CoVs, SARSCoV most likely originated in bats. For example, recombination probably occurred in bats before SARS in the event of the SARS outbreak in 2002-2003.
CoV was introduced from Yunnan into the province of Guangdong by infected civets or other infected mammals. The introduced SARS-CoV was subjected to rapid S and orf8 mutations and successfully distributed in market civets. Some of the strains suffered further S mutations after many separate spillovers to humans and became epidemics. Similarly, MERSr- CoVs outbreak in 2012 have been found in different bat species in many countries. Since large number of closely related coronaviruses are passed through several number of bat species, receptor usage plasticity and adaptive evultionary process is expected. Thus, interspecies transmission from bat reservoirs to animals and humans is anticipated [35-41]. Expectedly, high enetic similarities between SARS-CoV-2 and the sub-genus bat betacoronavirus Sarbecovirus was substantiated [42-47]. Even though, the means of transmission from natural reservoirs to humans still unclear, several investigations showed that pangolin’s virus may have given a partial spike gene, essential functional sites, to the new SARS-CoV-2 [48-50]. Evolutionary, many viruses have coexisted in/within their natural reservoirs for an exceptionally long time. However, continuous spillover of viruses from natural hosts to humans have occurred. This is basically because of human activities and behavior such as contemporary agricultural methods, invading natural habitats though urbanization and most importantly consuming exotic/wild animals. Thus, the most efficient means of preventing viral zoonosis is to sustain the barriers between natural reservoirs and human society and the implementation.
Integrative efforts by various disciplines to achieve optimum health for humans, animals and the environment, operating locally, nationally and internationally.
Coronavirus 2019 (COVID-19) disease is alternative name of the respiratory infectious disease due to extreme acute coronavirus respiratory syndrome (SARS-CoV-2) [51]. Although we are not Sure about the first identification date but the disease caused by the novel virus was officially first report in December 2019 in Wuhan, the capital of China's Hubei province. The novel infectious virus has since spread worldwide, resulting in the current pandemic of coronavirus 2019-20 [52,53].
More than 2.5 million cases have been registered across 185 countries as of 22 April 2020, resulting in more than 171,000 fatalities, and 71581 people have recovered [57].
Suspicion of infection, risk factors, but it is not recommended procedure for the routine identification [64,65].
Up to date April 22-2020, there is no specific treatment for COVID-19 [56,73], and research is ongoing for
specific medication and vaccine for this particular virus. Moreover, for the symptomatic patients, the firstline
prescription for paracetamol (acetaminophen) over ibuprofen is [74-76]. The FDA notes that there is
currently no proof that NSAIDs exacerbate the symptoms of COVID-19 [77]. and the use of non-steroidal
anti-inflammatory drugs (NSAIDs) such as ibuprofen for symptoms is not included in the WHO [78].
Previous SARS-CoV work is being used since both SARS-CoV and SARS-CoV-2 use the receptor of
ACE2 to enter human cells. For the Vaccination research there are three strategies being investigated. The
first, one aiming For the entire virus vaccine to be built. The aim of such a virus, be it inactive, to cause a
rapid immune response in the human body to the new infection of COVID-19. The second strategy, subunit
vaccines, aiming to generate a vaccine that stimulate the immune system to firm subunits of the virus. In
this The S-spike protein that helps the virus impose the ACE2 enzyme receptor is the research scientist's
priority. Nucleic acid vaccines (DNA or RNA vaccines, a modern vaccination producing technique) are
the third approach. For protection, any of the previous strategies must be tested, and effectivenes [79]. On
16 March 2020, the first clinical vaccine trial began in Seattle with four volunteers. The vaccine requires a
harmless genetic code copied from the disease-causing virus [80].
Furthermore, enhancement Antibody dependent has been proposed as a potential trial for vaccine development for COVID-19 [81].
The first detection of Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), or novel severe
acute respiratory syndrome coronavirus, was associated with a cluster of cases of acute respiratory disease
in Wuhan in three individuals with pneumonia [59]. All features of this novel virus occur in related to
coronaviruses in nature [83], with medium- sized (80-100nm) single positive strand RNA characteristics
enveloped particles surrounded by club- shaped projections.
Outside the human body, the virus is killed by household soap, which spurts its protective bubble [16]. SARS-CoV-2 is closely related to the original SARS-CoV [84]. The new mutative virus is a zoonotic origin, genetically clusters with the genus Betacoronavirus, in subgenus Sarbecovirus (lineage B) together with two bat-derived strains, with 96% homology at the whole genome near other bat coronavirus samples (BatCov RaTG13) [85]. Furthermore, In February 2020, Chinese researchers have found that there is only one amino acid discrepancy between pangolin viruses and human viruses in some parts of the genome sequences. The whole genome comparison to date shows that between pangolin coronavirus and SARS-CoV-2, which is still insufficient to prove pangolins to be the intermediate host, at most 92 percent of nucleic acid content was prevalent. The Chinese researcher discovered that there is only one variation between some parts of the amino acid, difference in certain parts of the genome sequences difference in certain parts of the genome sequences difference in certain parts of the genome sequences difference in certain parts of the genome sequences between the viruses from pangolins, and the one from humans. The, whole-genome contrast to date originate that at most 92% of nucleic acid material was common between pangolin coronavirus, and SARS-CoV-2, which is still insufficient to prove pangolins to be the intermediate host.
Epidemiology
The virus outbreak described as a pandemic with no clear peak incidence. The numbers are varying by region,
and over time and are predisposed by the size of testing, time since initial outbreak, and population features
such as age, sex, health healthcare quality and, treatment choices [86].
Based on Johns Hopkins University statistics ratio that reflecting death is 6.9% (176,926/2,560,504) as of 21 April 2020 [57]. This number varies by region [87].
Other procedures using the case fatality rate (CFR), that reflects the percent of diagnosed individuals who die from a disease, and the infection fatality rate (IFR), it is reflects the percentage of infected individuals (diagnosed, or undiagnosed) who die from a disease. The previous statistics approaches are not time bound, and follow a specific population from infection over case determination [88-91].
Bibliography

Hi!
We're here to answer your questions!
Send us a message via Whatsapp, and we'll reply the moment we're available!