Biography
Interests
Stavroula Koulocheri, A.
Department of Biological Chemistry, Medical School, National and Kapodistrian University of Athens, NKUA, Greece
*Correspondence to: Dr. Stavroula Koulocheri, A., Department of Biological Chemistry, Medical School, National and Kapodistrian University of Athens, NKUA, Greece.
Copyright © 2021 Dr. Stavroula Koulocheri, A. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Abstract
The ability of halogens to form non-covalent interactions, such as halogen bonding, and other polar interactions, in biomolecular structures was revealed recently. In this article, the concept of halogen bonding and the general background of σ-hole is presented, with special focus on its role in biomolecular sciences. Also, the geometry, the strength and its occurrence in biomolecules and in protein-ligand complexes is discussed. From the analysis of data, the potential use of halogen bonds is examined as molecular tools in the biomolecular recognition and the rational drug design.
Introduction
It is known that halogens are elements with high electronegativity and they function usually as electron
donors (nucleophiles) because of the concentrated electron density. Thus they can interact with electron
acceptors (electrophiles). Conventionally, halogens in ligands are used as electron-rich mainly, in non specific
interactions with biomolecules to the recognition of ligands by proteins.
However, there is another possibility that halogens can act as electron acceptors. The possibility of halogen atoms to function as electrophilic sites has been fully recognized only recently.
According to the definition proposed by the International Union of Pure and Applied Chemistry, the term halogen bond XB is used in attractive interactions where the halogen atom functions as electrophile and an electron-rich partner enters their positive region(s) [1].
It is denoted as R-X…Y. R-X is called halogen bond donor (electron acceptor), and X is any halogen atom i.e., I, Br, Cl, (rarely F), with an electrophilic (electron-poor) region. Y is the halogen bond acceptor functioning as nucleophile. Y, is an electron rich like O, N, S, or X-donor (Figure 1).
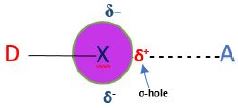
δ+ → electrophilic region
δ- → nucleophilic region
A=XB Acceptor X=I, Br, Cl D=XB Donor
In a typical halogen-bonding R-X…Y the interatomic distance tends to be less than the sum of the van der Waals (<ΣRvdW ) radii. Generally the angle R-X…Y tends to be close to 180° and the forces in halogen bonding are mainly of electrostatic nature including charge transfer, dispersion and polarization.
The σ-hole Model
The ability of halogens to form X-bonding can be explained by the development of the concept of “σ-holes” (early 2000’s) [2].
According to this model, the halogen atoms have the atomic structure s2 p2x p2y p1z -in atomic orbitals of the valence shell and the pz1 electron is responsible of the σ-bond to a carbon atom. This results in the depletion of electronic charge of the lobe of pz which is opposite the C-X σ-bond along the extension of the covalent bond formed by halogens. This leads to electron deficient hole. A region of positive electrostatic potential-a positive σ-hole is formed with a partial positive nuclear charge. The position of this region explains the XB directionality.
So, the electron density in covalently bound halogens through σ-bond is anisotropically distributed. Often, due to the anisotropic distribution of the electron density, halogen atoms show an amphoteric character. In this hole an electropositive cap appears and the four electrons in the px, py orbitals justify the electronegative ring, perpendicular to the σ-bond [2,3].
The electrophilic region is called “σ-hole” and is due to the anisotropic electron density distribution of halogens. Thus, a halogen at its σ-hole may act as an electron acceptor toward a Lewis base in a direction opposite to the covalent bond [1b]. This explains the linearity of the halogen bond. Because of the halogen character and the lack of additional bonds, the access to the σ-hole is easy sterically and electronically.
The size of the σ-hole (electropositive cap or crown) and the strength of the X-bond is dependent on the polarizability of the halogen and the negative inductive effect of groups covalently bonded to the halogen. With the increase of the size of the atom with F<Cl<Br<I and the consequent polarizability, the strength of X-bonds is increasing. F shows a weak σ-hole and is a weak halogen donor. π-electrons associating with aromatic amino residues and peptide carbonyl group can be X-bonds acceptors, although weaker as bases. Generally, the Lewis acid of the donor and the Lewis base of the acceptor determine the strength of X-bonding.
The electrophilic region can form attractive interactions with nucleophilic sites. Through such positive “σ-holes,” the atoms can interact attractively and directionally with negative regions as with the lone pairs of Lewis bases, anions, π electrons, etc.
Halogen bonding is a weak, electrostatically driven (encompassing polarization and dispersion), short range (<20kJ/mol), noncovalent interaction between an electrophilic region of a halogen and a nucleophile.
Halogen bonding in halogenated molecules seems to offer specificity in the recognition. This concept can be exploited at strategies for the rational design of new halogenated inhibitors against biomolecular targets (tool for biomolecular design) with the incorporation of X-bonds [4].
It is known that biomolecules generally do not contain halogens but are used to the resolution of X-ray structures of proteins and nucleic acids and are added to ligands [5].
Biomolecular Halogen Bonding (BXB) Donors and Acceptors
Inspecting biomolecular halogen bonding, the number of donors is small because halogen atoms are rare in
biomolecules. Usually halogens as BXB donors are found in proteins and polynucleotides technically for the
analysis of crystals during X-ray diffraction [6].
Also, a pair of Br-BXBs forces the DNA into a four-stranded Holliday junction (the four-stranded intermediate associated with genetic recombination). It is worth of note that halogenated amino acids in proteins, like chloro- and bromotyrosines derive from the action of oxidative halogenation by myeloperoxidases and eosinophil peroxidases, which serve as markers for asthma [7].
A very interesting class is the halogenated ligands acting as BXB donors. Numerous I…O contacts play important role in thyroid hormone recognition.
It is worth of mention the naturally occurring XB Donors, thyroxine-class of hormones, 3,5,3’-triiodo- L-thyronine (T3) and 3,5,3’5’-tetraiodo-L-thyronine (T4). The interactions and selectivity with the corresponding receptors are mediated through BXB (recognition of T3 by human thyroid hormone receptor) [8]. Also, structural analysis of protein transthyretin (TTR) transporting thyroxine hormones (T4) has shown the implication of X-bonds [9].
Thus, BXBs play roles in the recognition by receptors and transport proteins, and in the chemistry that converts between the two functional forms of the hormone.
X- bond donors include biomolecules as well as synthetic molecules (e.g., pharmaceuticals). A large group of BXB donor molecules concerns the inhibitors to various protein targets, with pharmacological interest.
Some non-steroidal anti-inflammatory drugs show BXB interactions with protein targets.
Because X atom can interact with both electron- rich and poor sites (Figure1) there is the possibility of halogen-water-hydrogen (XWH) bridges as it happens with diclofenac (2-[2-(2,6-dichloroanilino)phenyl] acetic acid ) a pain reliever (NSAID), H2O for simultaneous XB with diclofenac and HB with cytochrome P450 complex. It is difficult to exploit the XWH bridges in rational design, but XB is important in stabilizing conformations [10].
Also, triclosan, (2,4,4-trichloro-2-hydroxydiphenyl ether) is a broad spectrum antimicrobial agent which involves blocking of lipid biosynthesis in the microbe. Many halogenated pharmacological compounds are used as drugs or are tested in clinical trials.
It is known that protein kinases are targets for the treatment of viral infections, cardiovascular disease, cancer and inflammatory diseases. It seems that development of X bonding inhibitors is important with great usefulness [11].
There are many other examples showing that BXBs have the potentiality in drug research with focus on important targets. An example is the protein kinase CDK2 inhibitor 4,5,6,7-tetrabromo-benzotrazole [12].
BXB acceptors are considered Lewis bases similar to H-bond acceptors, and the ubiquitous water solvent. A usual acceptor base is the known carbonyl oxygen (C=O) of protein backbone and this group is the most frequent BXB acceptor in ligand complexes. It is known that polar amino acid residues in proteins expose side chain to water and preferentially form H-bonds. The aromatic side chains participate as BXB acceptors of the type C-X…π interactions. The bromines of some inhibitors of protein kinases seems to form π-BXBs with Phe side chains of CK2 and CDK2 offering a stabilizing interaction [11]. Regarding water solvent, in the presence of halogens waters can act as X-bond acceptors or as H-bond donors [13]. This depends on the orientation aligned or perpendicular to the σ-hole.
Geometry-Energy of BXB Bonds
The presence of a BXB appears as a short distance (shorter than the sum of the respective van der Waals
radii, ΣRvdW) between a halogen and a Lewis base atom. So every contact <ΣRvdW is a BXB [4,1b].
The nature of electronic characteristic of halogen and the Lewis base determines the great directionality of X-bonds, regarding the donor and acceptor. However, there are restrictions because of the crowded environment in proteins resulting in angular directionality of X-bonding interaction.
Strength can be varied (tunability) by modification of XB donor and acceptor. This is related with donor ability (I>Br>Cl>F) and the hybridization of C of C-X bond (sp>sp2>sp3) because of the difference in electronegativity of C. The negative inductive effect of groups on atom the XB donor is bound to (e.g., -CN, -NO2) can change the strength.
The X-bond is highly directional interaction following the model of σ-hole. Chemical complexity of biological systems leads to diverse set of interactions (less geometrically/structurally defined).
Examination of biomolecular structures in Protein Data Bank show angles approaching near linearity about 160º-165º for X-bond acceptor to the halogen contacts. Regarding the angle of the halogen toward the acceptor atom, there is an attraction of the σ-hole to the nonbonding electrons of the acceptor and the angle is about 120º [4,14].
However, there are exceptions relating with biological X-bonds because aromatic amino residues side chains and peptide bond involve delocalized π-electrons. Such π-X bonds show perpendicular direction to the aromatic ring. This occurs in the complexes of inhibitors to the protein kinases CDK2 and CK2 [11]. HIV1-reverse transcriptase inhibitor is a highly potent inhibitor (IC50 = 2nM) with a network of H-bond, π-π, and halogen bond [15].
It has to be mentioned that biological X-bonds are orthogonal interactions to H-bonds in case there is the carbonyl oxygen of a peptide bond as the same acceptor. So a X-bond is an additional interaction to existing H-bond of protein target. X-bonds, relating to H-bonds, are class of “weak” molecular interactions -with energies below ~12kcal/mol. X-bonds energies are in the range of from 1 to 5kcal/mol. The energies of X-bonds vary with geometry and structure [16].
Besides the relationship structure -energy of BXBs, the examination and elucidation of σ- hole bonding is necessary. The modeling of this weak interaction is of great importance. Computational modeling of the BXB, molecular design of BXBs as a tool, can help to the rational design of halogenated inhibitors against therapeutic targets and this field is the focus of many scientists [15,17]. Geometries and directionality of the BXBs may play an important role to the optimization efficacy of unhalogenated lead compounds [18]. Also there is the potential application of BXBs in biomolecular engineering. There are BXBs in DNA junctions which are, the essential building blocks for many biomolecular engineering applications [19]. Also, there are other applications in protein engineering [20].
Perspectives
The study of this weak interaction and the details of the structure and energies of BXBs is very important.
Applications of halogen bonds include crystal engineering, halogen bonding in biological settings, catalysis,
medicinal chemistry and rational drug design.
Even though the strength of this intermolecular force is near the range of thermal fluctuations, this interaction is associated with molecular folding and binding interactions. The contribution of each one of various interactions in a group of interactions usually is not the same. A weak interaction like halogen bond at an important binding site or at an active site of an enzyme may have a great effect in terms of specificity and function. Examining the affinity of a drug for its biological target and the transition from micromolar range to nanomolar range is very important. Tuning the strength of halogen bonds is of great importance during the lead optimization process. BXBs are important tools for rational design of halogenated compounds in drug development.
Generally, many BXBs are found in protein-ligand complexes. It is known that many drugs are halogenated. With the aid of docking programs there is the possibility and the challenge to make and design potent drugs exploiting the role that halogens play in drug design.
Affinity/specificity of these compounds is very important to target proteins.
BXBs are becoming well analyzed through database survey of this interaction in protein-ligand complexes. Computational and experimental studies associated with structure-energy relationships are increasing continually. X-bonding has been recognized by the biological experts, very important in its role in the binding specificity of halogenated inhibitors against protein targets. Halogen bonding should be used as a powerful tool, comparable with hydrogen bonding, to reinforce the binding affinity furthermore to regulate the binding selectivity. This non-covalent interaction can modify the molecular conformation. The unique properties of X-bonds, in cooperation with H-bonds, will allow BXBs to be exploited as a powerful molecular tool in drug discovery and biological design. Rational design of new and potent inhibitors against therapeutic targets through halogen bonding is very challenging for medicinal chemists and experts in this field. Halogen bonding, this non-covalent interaction has really a great potential in life sciences in the forthcoming years.
Bibliography

Hi!
We're here to answer your questions!
Send us a message via Whatsapp, and we'll reply the moment we're available!